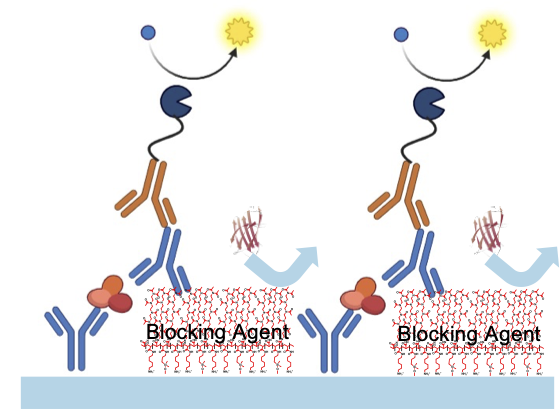
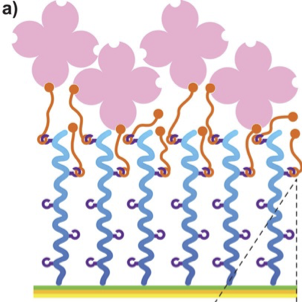
- Non-specific prevention of protein uptake
- Resistance against small-molecule drugs, DNA/RNA, cells and/or bacteria adhesion
- Hydrophilization reducing air-bubble formation
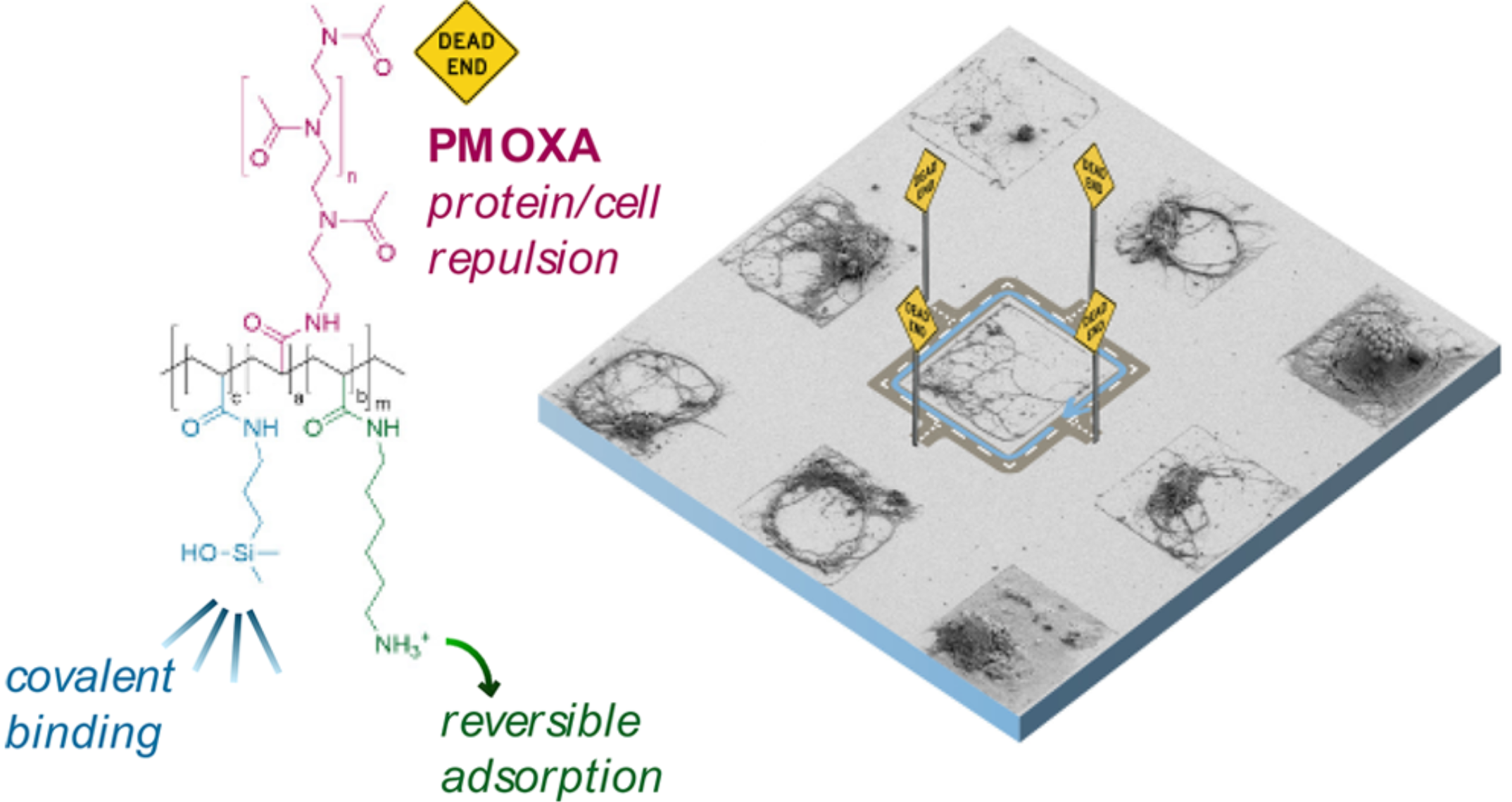
Poly-Oxazoline-based protein resistance coupled to specific binding.
Broad-band low-retention (non-fouling) monolayers prevent the adhesion of proteins, oligos, DNA/RNA or cells. SPAAC-clickable Azides (N3) allow for specific binding of bio-chemical groups.
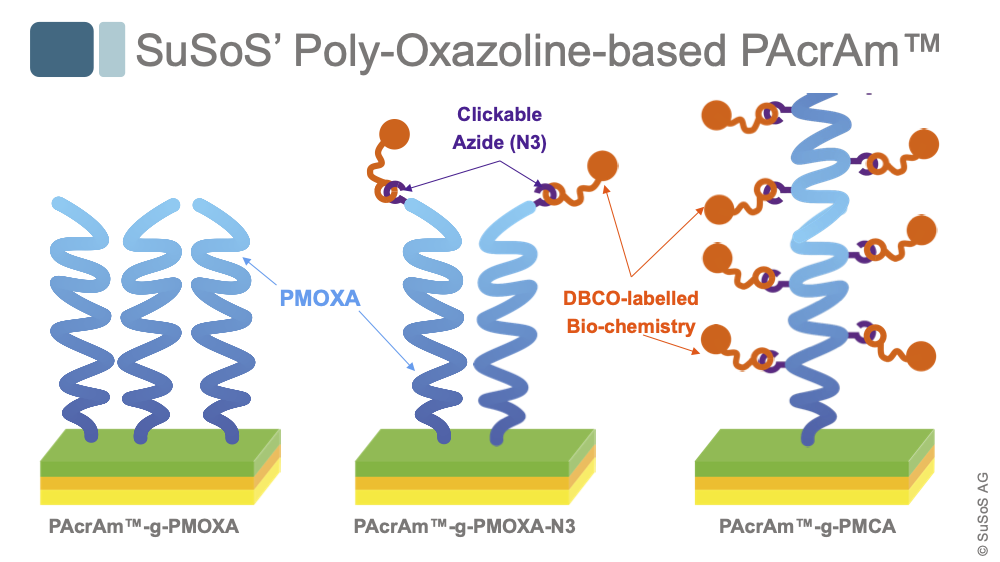
Off-the-shelf PAcrAm™ carry broad-band non-fouling PMOXA-based side chains that, while giving the surface a hydrophilic character, prevent the adhesion of bio-molecules and inhibit the formation of bubbles.
Thanks to SuSoS' proprietary chemistry, poly-oxazoline-based side chains are available with SPAAC-clickable azide (N3) groups.
We offer PMOXA-N3 with single functionalizations per PMOXA side-chain and PMOXA-co-N3-POXA (PMCA) multi-azide functionalizations per PMOXA side-chain.
PMOXA superiority
Contrarily to more traditional PEGs, PMOXA reportedly shows
- no oxidation
- high temperature reliability
- excellent UV stability
- superior non-fouling character for DNA/RNA oligomers or relatively "aggressive" cells like dendrites
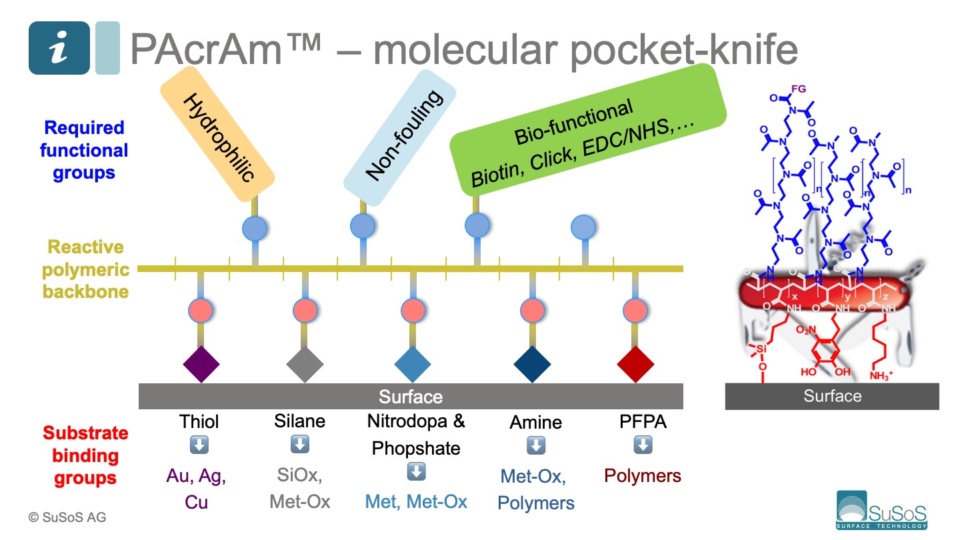
PAcrAm™ consistent polymer carpet monolayer.
PAcrAm™ consistently forms a stable, defect-free monolayer that irreversibly binds to specific substrates.
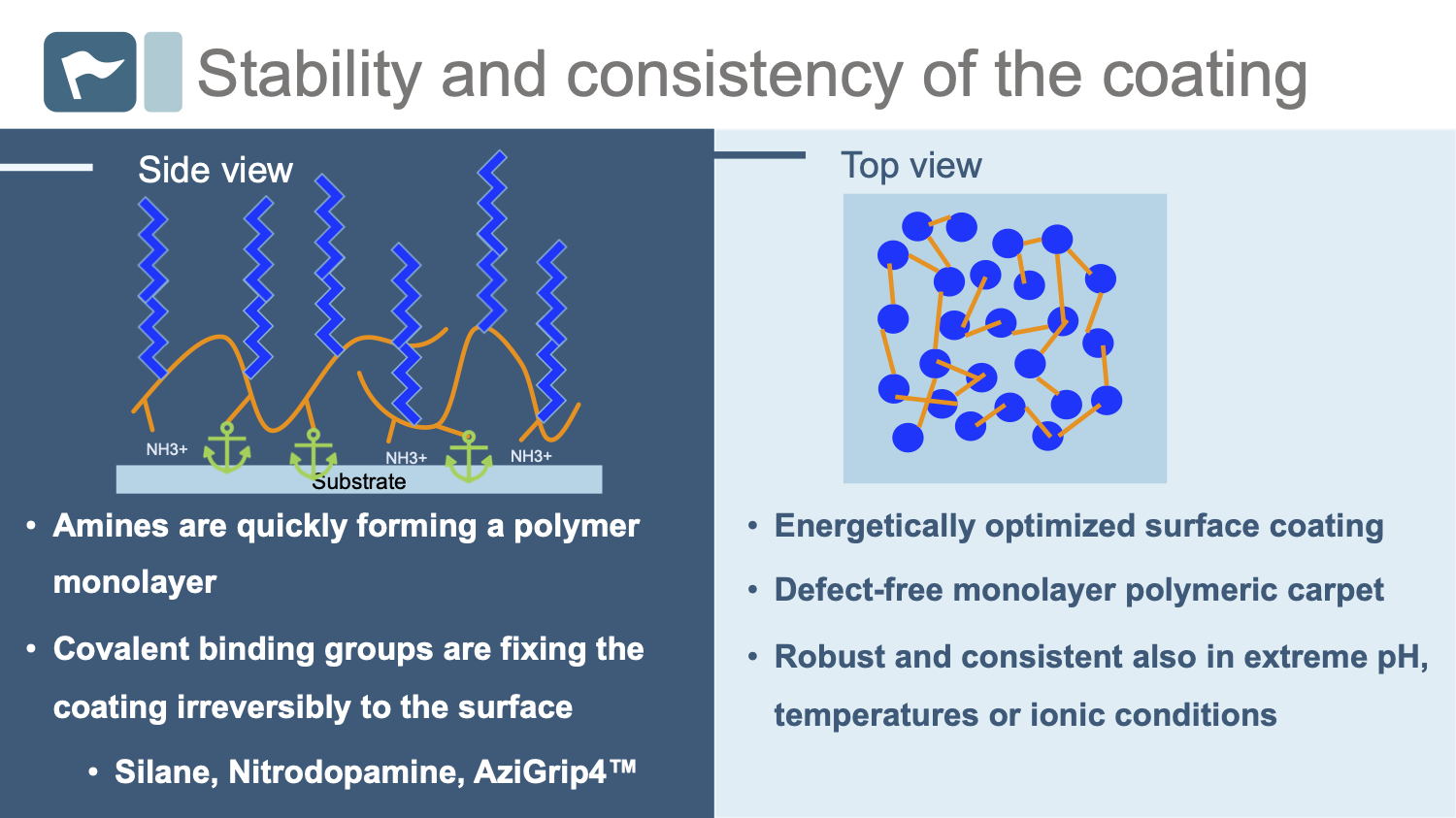
The elegant design of PAcrAm™ ensures a monolayer is quickly forming a continuum carpet on the surface thanks to instant electrostatic interactions through the amines in the polymer backbone.
The specific surface-reactive groups have then the time to covalently and irreversibly fix the energetically optimized carpet to the substrate.
Off-the-shelf PAcrAm™ backbones always positively-charged amine groups (NH3+) and specific surface-reactive
- silanes for glass, oxide and silicone (PDMS) substrates
- nitrodopamines for metals and metal oxide substrates
- AziGrip4™ for generic (non-reactive) polymer substrates
Select PAcrAm™ literature and applications
Providing a non-fouling, bio-functional coating on glass, titanium substrates and more
PacrAm™-based polymers carry strong covalent binding groups for different substrates (for example silicon or titanium oxides) for ultra-stable coatings by simple dip-and rinse protocols. Typical selective bio-functionalizations are biotin, RDG, NTA, SPAAC click and more…
- Serrano, A. et. al. Imparting Nonfouling Properties to Chemically Distinct Surfaces with a Single Adsorbing Polymer: A Multimodal Binding Approach. Macromol. Rapid Commun. 2016, 37 (7), 622–629. https://pubmed.ncbi.nlm.nih.gov/26858017/.
Mertgen, A. et. al. A Low-Fouling, Self-Assembled, Graft Co-Polymer and Covalent Surface Coating for Controlled Immobilization of Biologically Active Moieties. Appl. Surf. Sci. 2022, 584, 152525. https://doi.org/10.1016/j.apsusc.2022.152525.
PAcrAm™-g-(PMOXA,NH2,Si) prevents the adhesion of proteins to hollow AFM cantilevers for single cell manipulations, to inject biomolecules into cells or extract cell material for analysis from single cells. The coating greatly reduces adhesion of cell material or biomolecules to the cantilever walls and inside the microchannel.
- Li, M. et. al. FluidFM for Single-Cell Biophysics. Nano Res. 2021, 12 (1). https://doi.org/10.1007/s12274-021-3573-y – https://www.cytosurge.com
- Lüchtefeld, I. et al. Dissecting cell membrane tension dynamics and its effect on Piezo1-mediated cellular mechanosensitivity using force-controlled nanopipettes. Nat Methods 21, 1063–1073 (2024). https://doi.org/10.1038/s41592-024-02277-8
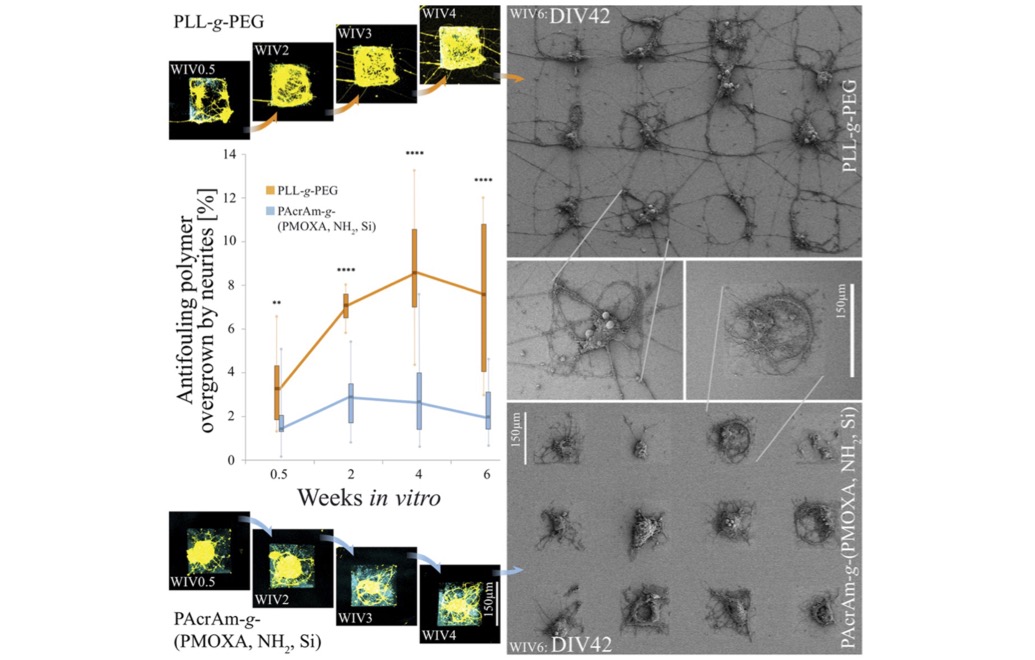
Selective long-term blocking of cell-adhesion and patterning
PacrAm™-g-(PMOXA, NH2, Si) has been the first surface coating that reliably prevents the formation of undesired connections between arranged nodes of primary neurons for up to 6 weeks in vitro.
- Weydert, S.et.al. Easy to Apply Polyoxazoline-Based Coating for Precise and Long-Term Control of Neural Patterns. Langmuir 2017, acs.langmuir.7b01437. https://doi.org/https://doi.org/10.1021/acs.langmuir.7b01437.
- Shahrokhtash, A. et. al. Microplate Format Protein Nanopatterning for High-Throughput Screening of Cellular Microenvironments. RXIV 2023, 1–24. https://doi.org/https://doi.org/10.1101/2023.11.19.567703.
Abdominal Imaging Window (AIW) implants coated with PAcrAm™-(PMOXA,NH2,Si) show an enhanced long-term visual clarity for intravital imaging of a mouse kidney.
- Sardella, D. et. al. Serial Intravital 2-Photon Microscopy and Analysis of the Kidney Using Upright Microscopes. Front. Physiol. 2023, 14 (April), 1–17. https://doi.org/10.3389/fphys.2023.1176409
Reducing loss of analyte molecules in POC devices
The PacrAm™-based coating used for AbioSCOPE™ sensor-chips reduces the loss of analyte due to nonspecific binding outside the sensing area leading to improved sensitivity and speed. https://www.abionic.com
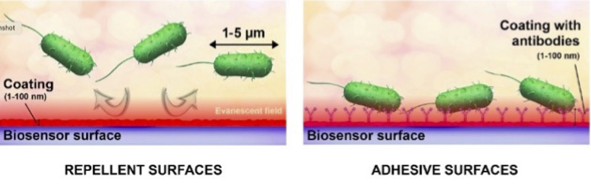
Bacterial resistance
PacrAm™-g-(PMOXA, NH2, Si) outperfoms state-of-the-art coatings such as PLL-g-PEG, albumin or other blocking strategies to reduce bacterial binding to surfaces.
- Farkas, E. et.al. Development and In-Depth Characterization of Bacteria Repellent and Bacteria Adhesive Antibody-Coated Surfaces Using Optical Waveguide Biosensing. Biosensors 2022, 12 (2). https://doi.org/10.3390/bios12020056.
Surface functionality to ultrasensitive molographic biosensors
PAcrAm™-g-(PEG-N3,NH2,ND) provides the initial coating to construct the molographic pattern, which is the core of the Ultrasensitive Biosensors. Azide-Alkyne click chemistry is used to attach photoactivatable groups to the sensor surface. Nitro-dopamine (ND) provides covalent binding to the Ta2O5 optical waveguide surface for stable coating performance https://www.lino-biotech.com
- Gatterdam, V et. al. Focal Molography Is a New Method for the in Situ Analysis of Molecular Interactions in Biological Samples. Nat. Nanotechnol. 2017, 37, 1502. https://doi.org/10.1038/nnano.2017.168
- Frutiger, A. et. al. Principles for Sensitive and Robust Biomolecular Interaction Analysis: The Limits of Detection and Resolution of Diffraction-Limited Focal Molography. Phys. Rev. Appl. 2019, 11 (1), S49. https://doi.org/10.1103/PhysRevApplied.11.014056
Blocking agent for ELISA-type assays
Thanks to its excellent non-fouling character, PAcrAm™-g-(PMOXA,NH2,Si) is outperforming standard blocking agents such as BSA, I-block, and PLL-g-PEG in antibody-based coatings for (bacterial) ELISA assays.
- Farkas, E. et.al. Development and In-Depth Characterization of Bacteria Repellent and Bacteria Adhesive Antibody-Coated Surfaces Using Optical Waveguide Biosensing. Biosensors 2022, 12 (2). https://doi.org/10.3390/bios12020056.
Increasing selective binding capacity up to a factor of 5 with PAcrAm™ PMCA
The multi-azide PAcrAmTM PMCA is shown to increase sensitivity compared to single-azide moieties by a factor of 3x-5x while maintaining the non-fouling properties typical of PMOXA- based coatings.
- , , Non-Fouling Multi-Azide Polyoxazoline Brush-co-Polymers for Sensing Applications. Adv. Mater. Interfaces 2024, 11, 2400322. https://doi.org/10.1002/admi.202400322
Thanks to azide-functionalized PAcrAm™, bioorthogonal tags at a submicron scale, can be used for arranging up to three distinct proteins,DNA, or peptides on large, fully passivated surfaces.
- Shahrokhtash A. and Sutherland. D.S., Smart Biointerfaces via Click Chemistry-Enabled Nanopatterning of Multiple Bioligands and DNA Force Sensors, ACS Applied Materials & Interfaces 2024 16 (17), 21534-21545. https://doi.org/10.1021/acsami.4c00831.
Minimize protein adsorption in solid state nanopores for single molecule detection
Covalently-attached PAcrAm™-g-(PMOXA,NH2,Si) to the inside of solid state nanopores
- minimize nonspecific surface interactions between biomolecules and the pore walls favoring free translocation of proteins,
- enable repeated and prolonged experiments of protein translocation without clogging and baseline shifts,
- eliminate the need for stringent cleaning protocols of nanopores between experiments,
- enable estimates of volume and length-to-diameter ratio of untethered, unlabeled proteins that are in close agreement with reference values.
Awasthi, S. et. al. Polymer Coatings to Minimize Protein Adsorption in Solid‐State Nanopores. Small Methods 2020, 2000177. https://doi.org/10.1002/smtd.202000177
Aramesh, M. et. al. Localized Detection of Ions and Biomolecules with a Force-Controlled Scanning Nanopore Microscope. Nat. Nanotechnol. 2019, 14 (8), 791–798. https://doi.org/10.1038/s41565-019-0493-z
Schlotter, T. et. al. Force-Controlled Formation of Dynamic Nanopores for Single-Biomolecule Sensing and Single-Cell Secretomics. ACS Nano 2020, 14 (10), 12993–13003. https://doi.org/10.1021/acsnano.0c04281
Schlotter, T. et. al. Aptamer-Functionalized Interface Nanopores Enable Amino Acid-Specific Single-Molecule Proteomics. Preprint 2023, 1–21. https://doi.org/10.21203/rs.3.rs-3015491/v1
Custom– PAcrAm™
PAcrAm™ can be synthesized virtually to carry any functionalization. Talk to us to find out how we can help you.
The PAcrAm™ technology is covered by international patents controlled by SuSoS. PAcrAm™ is SuSoS trade mark. All rights reserved.